Introduction
In the previous post I gave a very brief introduction to ABC, including a simple example for inferring the parameters of a Markov process given some time series observations. Towards the end of the post I observed that there were (at least!) two potential problems with scaling up the simple approach described, one relating to the dimension of the data and the other relating to the dimension of the parameter space. Before moving on to the (to me, more interesting) problem of the dimension of the parameter space, I will briefly discuss the data dimension problem in this post, and provide a couple of references for further reading.
Summary stats
Recall that the simple rejection sampling approach to ABC involves first sampling a candidate parameter 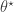 from the prior and then sampling a corresponding data set
from the prior and then sampling a corresponding data set 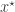 from the model. This simulated data set is compared with the true data
from the model. This simulated data set is compared with the true data  using some (pseudo-)norm,
using some (pseudo-)norm,  , and accepting
, and accepting 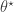 if the simulated data set is sufficiently close to the true data,
if the simulated data set is sufficiently close to the true data,  . It should be clear that if we are using a proper norm then as
. It should be clear that if we are using a proper norm then as  tends to zero the distribution of the accepted values tends to the desired posterior distribution of the parameters given the data.
tends to zero the distribution of the accepted values tends to the desired posterior distribution of the parameters given the data.
However, smaller choices of  will lead to higher rejection rates. This will be a particular problem in the context of high-dimensional
will lead to higher rejection rates. This will be a particular problem in the context of high-dimensional  , where it is often unrealistic to expect a close match between all components of
, where it is often unrealistic to expect a close match between all components of  and the simulated data
and the simulated data 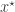 , even for a good choice of
, even for a good choice of 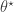 . In this case, it makes more sense to look for good agreement between particular aspects of
. In this case, it makes more sense to look for good agreement between particular aspects of  , such as the mean, or variance, or auto-correlation, depending on the exact problem and context. If we can find a finite set of sufficient statistics,
, such as the mean, or variance, or auto-correlation, depending on the exact problem and context. If we can find a finite set of sufficient statistics,  for
for 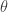 , then it should be clear that replacing the acceptance criterion with
, then it should be clear that replacing the acceptance criterion with  will also lead to a scheme tending to the true posterior as
will also lead to a scheme tending to the true posterior as  tends to zero (assuming a proper norm on the space of sufficient statistics), and will typically be better than the naive method, since the sufficient statistics will be of lower dimension and less “noisy” that the raw data, leading to higher acceptance rates with no loss of information.
tends to zero (assuming a proper norm on the space of sufficient statistics), and will typically be better than the naive method, since the sufficient statistics will be of lower dimension and less “noisy” that the raw data, leading to higher acceptance rates with no loss of information.
Unfortunately for most problems of practical interest it is not possible to find low-dimensional sufficient statistics, and so people in practice use domain knowledge and heuristics to come up with a set of summary statistics,  which they hope will closely approximate sufficient statistics. There is still a question as to how these statistics should be weighted or transformed to give a particular norm. This can be done using theory or heuristics, and some relevant references for this problem are given at the end of the post.
which they hope will closely approximate sufficient statistics. There is still a question as to how these statistics should be weighted or transformed to give a particular norm. This can be done using theory or heuristics, and some relevant references for this problem are given at the end of the post.
Implementation in R
Let’s now look at the problem from the previous post. Here, instead of directly computing the Euclidean distance between the real and simulated data, we will look at the Euclidean distance between some (normalised) summary statistics. First we will load some packages and set some parameters.
require(smfsb)
require(parallel)
options(mc.cores=4)
data(LVdata)
N=1e7
bs=1e5
batches=N/bs
message(paste("N =",N," | bs =",bs," | batches =",batches))
Next we will define some summary stats for a univariate time series – the mean, the (log) variance, and the first two auto-correlations.
ssinit <- function(vec)
{
ac23=as.vector(acf(vec,lag.max=2,plot=FALSE)$acf)[2:3]
c(mean(vec),log(var(vec)+1),ac23)
}
Once we have this, we can define some stats for a bivariate time series by combining the stats for the two component series, along with the cross-correlation between them.
ssi <- function(ts)
{
c(ssinit(ts[,1]),ssinit(ts[,2]),cor(ts[,1],ts[,2]))
}
This gives a set of summary stats, but these individual statistics are potentially on very different scales. They can be transformed and re-weighted in a variety of ways, usually on the basis of a pilot run which gives some information about the distribution of the summary stats. Here we will do the simplest possible thing, which is to normalise the variance of the stats on the basis of a pilot run. This is not at all optimal – see the references at the end of the post for a description of better methods.
message("Batch 0: Pilot run batch")
prior=cbind(th1=exp(runif(bs,-6,2)),th2=exp(runif(bs,-6,2)),th3=exp(runif(bs,-6,2)))
rows=lapply(1:bs,function(i){prior[i,]})
samples=mclapply(rows,function(th){simTs(c(50,100),0,30,2,stepLVc,th)})
sumstats=mclapply(samples,ssi)
sds=apply(sapply(sumstats,c),1,sd)
print(sds)
# now define a standardised distance
ss<-function(ts)
{
ssi(ts)/sds
}
ss0=ss(LVperfect)
distance <- function(ts)
{
diff=ss(ts)-ss0
sum(diff*diff)
}
Now we have a normalised distance function defined, we can proceed exactly as before to obtain an ABC posterior via rejection sampling.
post=NULL
for (i in 1:batches) {
message(paste("batch",i,"of",batches))
prior=cbind(th1=exp(runif(bs,-6,2)),th2=exp(runif(bs,-6,2)),th3=exp(runif(bs,-6,2)))
rows=lapply(1:bs,function(i){prior[i,]})
samples=mclapply(rows,function(th){simTs(c(50,100),0,30,2,stepLVc,th)})
dist=mclapply(samples,distance)
dist=sapply(dist,c)
cutoff=quantile(dist,1000/N,na.rm=TRUE)
post=rbind(post,prior[dist<cutoff,])
}
message(paste("Finished. Kept",dim(post)[1],"simulations"))
Having obtained the posterior, we can use the following code to plot the results.
th=c(th1 = 1, th2 = 0.005, th3 = 0.6)
op=par(mfrow=c(2,3))
for (i in 1:3) {
hist(post[,i],30,col=5,main=paste("Posterior for theta[",i,"]",sep=""))
abline(v=th[i],lwd=2,col=2)
}
for (i in 1:3) {
hist(log(post[,i]),30,col=5,main=paste("Posterior for log(theta[",i,"])",sep=""))
abline(v=log(th[i]),lwd=2,col=2)
}
par(op)
This gives the plot shown below. From this we can see that the ABC posterior obtained here is very similar to that obtained in the previous post using the full data. Here the dimension reduction is not that great – reducing from 32 data points to 9 summary statistics – and so the improvement in performance is not that noticable. But in higher dimensional problems reducing the dimension of the data is practically essential.

Summary and References
As before, I recommend the wikipedia article on approximate Bayesian computation for further information and a comprehensive set of references for further reading. Here I just want to highlight two references particularly relevant to the issue of summary statistics. It is quite difficult to give much practical advice on how to construct good summary statistics, but how to transform a set of summary stats in a “good” way is a problem that is reasonably well understood. In this post I did something rather naive (normalising the variance), but the following two papers describe much better approaches.
I still haven’t addressed the issue of a high-dimensional parameter space – that will be the topic of a subsequent post.
The complete R script
require(smfsb)
require(parallel)
options(mc.cores=4)
data(LVdata)
N=1e6
bs=1e5
batches=N/bs
message(paste("N =",N," | bs =",bs," | batches =",batches))
ssinit <- function(vec)
{
ac23=as.vector(acf(vec,lag.max=2,plot=FALSE)$acf)[2:3]
c(mean(vec),log(var(vec)+1),ac23)
}
ssi <- function(ts)
{
c(ssinit(ts[,1]),ssinit(ts[,2]),cor(ts[,1],ts[,2]))
}
message("Batch 0: Pilot run batch")
prior=cbind(th1=exp(runif(bs,-6,2)),th2=exp(runif(bs,-6,2)),th3=exp(runif(bs,-6,2)))
rows=lapply(1:bs,function(i){prior[i,]})
samples=mclapply(rows,function(th){simTs(c(50,100),0,30,2,stepLVc,th)})
sumstats=mclapply(samples,ssi)
sds=apply(sapply(sumstats,c),1,sd)
print(sds)
# now define a standardised distance
ss<-function(ts)
{
ssi(ts)/sds
}
ss0=ss(LVperfect)
distance <- function(ts)
{
diff=ss(ts)-ss0
sum(diff*diff)
}
post=NULL
for (i in 1:batches) {
message(paste("batch",i,"of",batches))
prior=cbind(th1=exp(runif(bs,-6,2)),th2=exp(runif(bs,-6,2)),th3=exp(runif(bs,-6,2)))
rows=lapply(1:bs,function(i){prior[i,]})
samples=mclapply(rows,function(th){simTs(c(50,100),0,30,2,stepLVc,th)})
dist=mclapply(samples,distance)
dist=sapply(dist,c)
cutoff=quantile(dist,1000/N,na.rm=TRUE)
post=rbind(post,prior[dist<cutoff,])
}
message(paste("Finished. Kept",dim(post)[1],"simulations"))
# plot the results
th=c(th1 = 1, th2 = 0.005, th3 = 0.6)
op=par(mfrow=c(2,3))
for (i in 1:3) {
hist(post[,i],30,col=5,main=paste("Posterior for theta[",i,"]",sep=""))
abline(v=th[i],lwd=2,col=2)
}
for (i in 1:3) {
hist(log(post[,i]),30,col=5,main=paste("Posterior for log(theta[",i,"])",sep=""))
abline(v=log(th[i]),lwd=2,col=2)
}
par(op)